Assumptions of linear regression, residuals, Q-Q plot

Residual Analysis in Linear Regression
Assumptions in Linear regression are about residuals. Let’s learn about residuals and assumptions in linear regression about residuals.
Residuals:
Residuals in Linear Regression are the difference between the actual value and the predicted value.

How is the predicted value calculated?
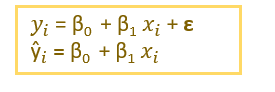
ε → Residuals or Error term.
Assumptions in Linear Regression are about residuals:
- Residuals should be independent of each other.
- Residuals should have constant variance.
- The expected value or mean of the residuals should be zero. E[ε]=0
- Residuals should follow a normal distribution
Residual Plots
- Residual vs target variable
- Residual vs predicted variable
- Distplot of residuals
Scenario 1: All assumptions are satisfied
Example: I have taken a simple dataset.

x- independent variable, y-target variable.
Before building a linear regression model, let’s check scatterplot,regplot, and heatmap.
df=pd.read_csv("ex1.csv")
Scatterplot
sns.scatterplot(df['x'],df['y'],color='darkorange')

regplot
sns.regplot(df['x'],df['y'],color='darkorange')

df.corr()

Building model and calculating residuals
import statsmodels.api as sm
X_train_sm = sm.add_constant(X)
fit1 = sm.OLS(y, X_train_sm).fit()
#Calculating y_predict and residuals
y_predict=fit1.predict(x_train_sm)
residual=fit1.resid
Assumption 1: Residuals are independent of each other.
Assumption 2: Residuals should have a constant variance.
To check these assumptions, we have to plot residual plot [y_predict vs residuals’]
sns.residplot(y_predict,residual,color='darkorange')
plt.xlabel("y_predict")
plt.ylabel("Residuals")
plt.title("Residual Plot")
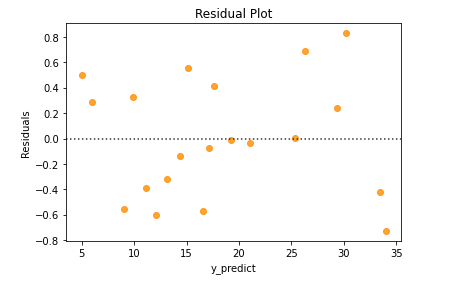
From the above residual plot, we could infer that the residuals didn’t form any pattern. So, the residuals are independent of each other.
And also, the residuals have constant variance. Variance doesn’t seem to increase/decrease constantly with the y_predict value.
Assumption 3: Residuals are normally distributed
To check whether the residuals are normally distributed from Q-Q plot,distplot
- Q-Q plot → quantile-quantile plot
If the residuals are normally distributed, then the Q-Q plot of residuals will be a straight line
from scipy import stats
import statsmodels.api as sm
residual=fit1.resid
probplot=sm.ProbPlot(residual,stats.norm,fit=True)
fig=probplot.qqplot(line='45')
plt.title('qqplot')

2. distplot
sns.distplot(residual,color='darkorange')

Scenario 2: Residuals are not independent of each other
I have created a small data set that contains x and y and y = x² with some noise added to it.
y →target variable
x →independent variable

- Scatterplot
df1=pd.read_csv("ex2.csv")
sns.scatterplot(df1['x'],df1['y'],color='darkorange')
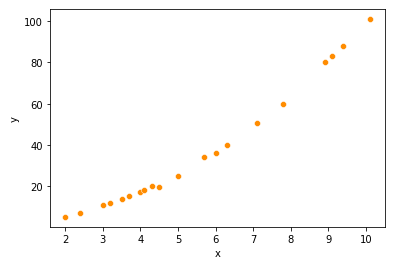
- Regplot
sns.regplot(df1['x'],df1['y'],color='darkorange')
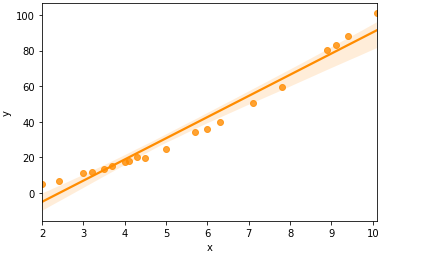
Correlation value

Building Model and calculating y_predict and residuals
X1 = np.array(df1['x']).reshape(-1,1) # predictor variable
y1 = np.array(df1['y']).reshape(-1,1) # response variable
import statsmodels.api as sm
x1_train_sm = sm.add_constant(X1)
fit2 = sm.OLS(y1, x1_train_sm).fit()
y1_predict=fit2.predict(x1_train_sm)
residual_1=fit2.resid
Residual plot
sns.residplot(y1_predict,residual_1,color='darkorange')
plt.xlabel("y1_predict")
plt.ylabel("Residuals")
plt.title("Residual Plot")

From the residual plot, we could see that the residuals follow a pattern. They are dependent on each other. Non-linearity is present in the data.
Since the residuals are dependent on each other, we can now build a slightly different model.
Let’s build a degree 2 polynomial model
from sklearn.preprocessing import PolynomialFeatures
polynomial_features= PolynomialFeatures(degree=2)
xp = polynomial_features.fit_transform(X1)
xp_train=sm.add_constant(xp)
fit_p= sm.OLS(y,xp_train).fit()
yp_predict=fit_p.predict(xp_train)
residual_p=fit_p.resid
Let’s check the residual plot for the new model
sns.residplot(yp_predict,residual_p,color='darkorange')
plt.xlabel("yp_predict")
plt.ylabel("Residuals")
plt.title("Residual Plot")

Now, the residuals are independent of each other.
Scenario 3: Residuals doesn’t have constant variance
If the residuals don’t have constant variance, we can try transforming independent variables (log-transformation,box-cox transformation)
If the residuals don’t have constant variance, we can infer it from the residual plot. If we get a residual plot like the one mentioned below, it means that residuals don’t have constant variance. Here, the residuals spread is not constant.

Conclusion:
In this article, I have covered residuals, the assumptions of residuals in linear regression, and plots to check the assumptions of residuals.
Thanks for reading!
If you like to read more of my tutorials on Python and Data Science,
follow me on Medium, Twitter
https://indhumathychelliah.medium.com/membership
Make a one-time donation
Make a monthly donation
Make a yearly donation
Choose an amount
Or enter a custom amount
Your contribution is appreciated.
Your contribution is appreciated.
Your contribution is appreciated.